Last time I wrote I gave a brief introduction to the Game of Life and a very simple Python implementation for visualizing it. I will freely admit that was a teaser post; this post gets into the real meat of the topic with an overview of the HashLife algorithm and a much more interesting implementation.
Introduction
This entry has taken me an embarrassingly long time to post. As is my habit, I wrote the code and 90% of the post, and then left it for months and months. Whoops!
If you haven’t played with a Game of Life viewer before they are legitimately fun to toy around with - I encourage you to check this one out (code is here). Since the last version everything is much improved. The viewer supports a larger set of controls (see the README for details) and basic file reading is implemented so it’s possible to try new starting patterns on the fly. And, as promised, I’ve implemented the HashLife algorithm to massively speed up iterations, so enormous patterns billions of generations forward are easily within your reach.
Algorithm
HashLife is a simple yet interesting algorithm. Invented in 1984 by Bill Gosper (of Gosper glider gun fame), it exploits repeated patterns to dramatically cut down the work required to support large patterns over vast numbers of iterations. Between the Wikipedia page and the enigmatically named “An Algorithm for Compressing Space and Time” in Dr. Dobb’s Journal I think it’s decently well explained, but it took me a couple read-throughs to really wrap my head around so I’m going to try to give an overview of the key insights it utilizes.
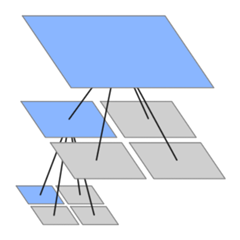
At it’s heart, HashLife is built around the concept of a quadtree. If you’re unfamiliar with it, a quadtree takes a square region and breaks it into four quadrants, each a quarter the size of the original. Each quadrant is further broken down into quadrants of its own, and on down. At the bottom, in squares of some minimum size like 2x2, actual points are stored. This structure is usually used to make spatial queries like “what points intersect this bounding box” efficient, but in this case two other properties are taken advantage of. First, nodes at any level are uniquely defined by the points within their region, which means duplicated regions can be backed by the same node in memory. For the Game of Life, where there are repeated patterns and empty regions galore, this can drastically reduce the space required. Second, in the Game of Life a square of (n)x(n) points fully dictates the inner (n-2)x(n-2) core one generation forward, the inner (n/2)x(n/2) core n/4 generations forward, irrespective of what cells are adjacent to it. So the future core of a node can be calculated once and will apply at any future point in time, anywhere in the tree.
 Together these properties allow for ridiculous speedups. Hashing and sharing nodes drastically reduces the space requirements, with exponentially more sharing the further down the tree you go. There are only 16 possible leaf nodes, after all! From this, calculating the future core for a node requires exponentially less time than a naïve implementation would. It can be done by recursively calculating the inner core of smaller nodes, where the better caching comes into play, and then combining them together into a new node. You might be wondering if the gains from caching are lost to the increasing difficulty of determining which nodes are equal, but with a couple careful invariants we actually get that for free. First, nodes must be immutable - this one’s pretty straightforward. Second, nodes must be unique at all times. This forces us to build the tree from the bottom up, but then checking if a new node duplicates an existing one is simply a matter of checking if there are any existing nodes that point to the same set of quadrants in the same order, a problem that hash tables trivially solve.
Together these properties allow for ridiculous speedups. Hashing and sharing nodes drastically reduces the space requirements, with exponentially more sharing the further down the tree you go. There are only 16 possible leaf nodes, after all! From this, calculating the future core for a node requires exponentially less time than a naïve implementation would. It can be done by recursively calculating the inner core of smaller nodes, where the better caching comes into play, and then combining them together into a new node. You might be wondering if the gains from caching are lost to the increasing difficulty of determining which nodes are equal, but with a couple careful invariants we actually get that for free. First, nodes must be immutable - this one’s pretty straightforward. Second, nodes must be unique at all times. This forces us to build the tree from the bottom up, but then checking if a new node duplicates an existing one is simply a matter of checking if there are any existing nodes that point to the same set of quadrants in the same order, a problem that hash tables trivially solve.
def __hash__(self): # Hash is dependent on cells only, not e.g. _next. # Required for Canonical(), so cannot be simply the id of the current # object (which would otherwise work). return hash((id(self._nw), id(self._ne), id(self._sw), id(self._se))) def __eq__(self, other): """Are two nodes equal? Doesn't take caching _next into account.""" if id(self) == id(other): return True return (id(self._nw) == id(other._nw) and id(self._ne) == id(other._ne) and id(self._sw) == id(other._sw) and id(self._se) == id(other._se))
Implementation
As before, the code I’ve written is for Python 2.6 and makes use of PyGame, although neither dependency is terribly sticky. The code lives in a repository on github, and I welcome any contributions you care to make. As the code here is complicated enough to be almost guaranteed a bug or two, there is a basic set of unit tests in life_test.py and the code itself is liberally sprinkled with asserts. Incidentally, removing the asserts nets a 20% performance gain (as measured by the time it takes to run the ‘PerformanceTest’ unit test), although I find the development time saved by having them is easily worth keeping them in forever. As noted later, the performance of the implementation isn’t all that important anyways. Which is a good thing, since I coded it in Python!
A comment on rewrites: during the transition from version 1 - a simple brute force algorithm - to version 2 - the Node class that implements HashLife - I had both algorithms implemented in parallel for a while. This let me have every second frame rendered by the old algorithm so I could ensure that at different times and different render speeds that the algorithms were coming up with the same results. I’ve seen this pattern used at work for migrating to replacement systems and it’s very much worth the extra glue code you have to write or the confidence it gives. John Carmack recently wrote about parallel implementations on his own blog, if you want to hear more on the topic.
Performance
The performance is hard to objectively detail for an algorithm like this. For example, it takes ~1 second to generate the billionth generation of the backrake 3 pattern, which has around 300,000,000 live cells; it takes ~2 seconds to generate the quintillionth generation with 3x10^17 live cells. But this is a perfect patterns to showcase HashLife - a simple spaceship traveling in a straight line, generating a steady stream of gliders. In comparison, a chaotic pattern like Acorn takes almost 25 seconds to generate just 5000 generations with at most 1057 alive at any time. As it stands the properties of the algorithm drastically outweigh the peculiarities of the implementation for anything I care to do. Although I must say, if you want to compare it to another implementation in an apples to apples comparison I’d love to hear the numbers you get.
As always, I’d love to hear what you think!
 I'm Eric Burnett, a software engineer at Google. I'm interested in studying what makes programmers tick, and writing whatever code strikes my fancy. Read my
I'm Eric Burnett, a software engineer at Google. I'm interested in studying what makes programmers tick, and writing whatever code strikes my fancy. Read my 